Phylogenetic Stochastic Mapping without Matrix Exponentiation
Jan Irvahn, @vminin
Phylogenetic stochastic mapping is a method for reconstructing the history of trait changes on a phylogenetic tree relating species/organisms carrying the trait. State-of-the-art methods assume that the trait evolves according to a continuous-time Markov chain (CTMC) and work well for small state spaces. The computations slow down considerably for larger state spaces (e.g. space of codons), because current methodology relies on exponentiating CTMC infinitesimal rate matrices – an operation whose computational complexity grows as the size of the CTMC state space cubed. In this work, we introduce a new approach, based on a CTMC technique called uniformization, that does not use matrix exponentiation for phylogenetic stochastic mapping. Our method is based on a new Markov chain Monte Carlo (MCMC) algorithm that targets the distribution of trait histories conditional on the trait data observed at the tips of the tree. The computational complexity of our MCMC method grows as the size of the CTMC state space squared. Moreover, in contrast to competing matrix exponentiation methods, if the rate matrix is sparse, we can leverage this sparsity and increase the computational efficiency of our algorithm further. Using simulated data, we illustrate advantages of our MCMC algorithm and investigate how large the state space needs to be for our method to outperform matrix exponentiation approaches. We show that even on the moderately large state space of codons our MCMC method can be significantly faster than currently used matrix exponentiation methods.
[Paper] Phylogenetic Stochastic Mapping without Matrix Exponentiation


Thanks for highlighting the paper, @ematsen! Jan and I would like to incorporate this approach into a larger MCMC, where we could estimate at least the substitution matrix and branch lengths (more ambitious plans include phylogeny estimation as well, but we have only preliminary ideas). We are on the look out for collaborators!

By the way, @ematsen labeled the paper as “traits,” but the method works for sequences as well. For example, it should be possible to “plug in” our new MCMC sampler into PhyloBayes machinery that deals with complex models of protein evolution.

@vminin, I’ve been working on a general modeling framework for stochastic mapping and large state spaces in RevBayes and am planning to add support for Rao-Teh uniformization in the future. Perhaps you will welcome a question or two as they arise?
I’ve also been toying with the idea of joint inference of stochastic mappings and phylogeny using MCMC, which I imagine will need this sort of rapid simulation for efficient joint proposals on topology and mappings (e.g. using delayed-rejection). It’d be fun to work out the details if you and Jan are interested.
(I was visiting Jeff Thorne at NCSU in Fall 2013 when you so happened to visit to chat about your progress on this work. Really excited to see it in print.)

@mlandis, of course I remember chatting with you at NCSU! Now I feel bad for missing and not citing your SysBio (2013) paper. I hope you didn’t have it posted as a preprint even earlier, because this would be even more embarrassing.
Jan and I would be happy to answer questions and share our ideas about future directions. Shoot me an email when you have a chance.
Have you looked at PhyloBayes source? I never did, so I could never quite understand whether it does or does not sample phylogenies and stochastic mappings jointly…

I noticed this paper last night on the arxiv mailing and was intrigued because we have been toying a little with the idea of avoiding matrix exponentials in the context of our work on Lie Markov models. I have no idea what uniformization is, so I’m sort of posting here as a place holder to chase up what this is all about. For me the interest is that, for models with the right algebraic properties, we can parametrize substitution probabilities directly (without using rate matrices or matrix exps), and then, after parameter estimation, calculate what the appropriate rate parameters would have been (via a matrix log, or even directly via explicit formulas for particular models).

Dear Authors, I think that an explanation of what a virtual jump was would be helpful. I struggled a bit getting a toe-hold in understanding this paper so I’m adding a little explanation for other future readers. I may be totally wrong here, but hopefully I will get corrected.
Jumps come in two flavors: real and virtual. The only difference is that virtual jumps don’t change state. State transitions are only allowed at jump positions. The MCMC used to sample jump histories only allows changes of real jumps through introduction of the jump as a virtual jump.
Here is a figure giving an example step through the MCMC.

Note that edge numbering is (for some confusing reason) only annotated in plot 3. Those numbers have nothing to do with the substitution process. The figure explains this process as follows:
-
Here is a substitution history which is augmented by the presence of virtual jumps. Again, we are just seeing these virtual jumps as jumps that don’t change state, so this is just a standard sampled history of a trait on a tree where self-transitions are allowed.
-
We go through the tree, sampling states conditioned on the data in a way such that we are only allowed to change state at the jump positions (real or virtual). The jumps that don’t change state become virtual, and the inverse.
-
and 4. Resample the virtual jumps, which will then become opportunities to change state in the next iteration.
Did I get it right, @vminin?

No, I’m not too familiar with PhyloBayes’ code either. It looks like PhyloBayes::Move implements a variety of topology proposals, but it’s not clear to me whether the likelihoods for these proposals are computed using matrix exponentiation or mapped histories. Maybe @nicolas_lartill could shed light on this?

My guess is that PhyloBayes updates topologies with the regular likelihood, computed with the help of matrix exponentiation, and then draws a realization of the substation histories when needed.
Since Jan and I have moved from Monte Carlo to MCMC this strategy will not fly anymore, and one would have to update topologies and (augmented) substitution histories jointly.

You got it, @ematsen! Yes, we need to clarify that self-transitions and virtual jumps are the same things. I think Jan had this clarification somewhere and it was me who accidentally removed during editing. Thanks, our paper hasn’t been assigned and AE yet, but we already are getting useful peer-reveiw!

PhyloBayes does not sample phylogenies jointly with substitution mappings. It is essentially a two-phase system: (1) sample a substitution history and update continous parameters conditional on this mapping (2) discard mapping, go back to integrated likelihood mode (using pruning) and update topology.
Incidentally, I would like to point out that PhyloBayes never calculates matrix exponentials.
the trick is this: when propagating conditional likelihoods over the tree, you need to do something like:
L_up = exp( t Q) L_down = P exp( t D) P^{-1} L_down
where L_up and L_down are the conditional likelihood vectors, D is the diagonal matrix containing the eigenvalues and P the eigenvector matrix.
now, if you essentially do: L_up = P [ exp( t D) [ P^{-1} L_down] ]
instead of L_up = [P [ exp( t D) P^{-1}] L_down
then you do not need to compute the exponential. On the other hand, you have two matrix-vector products to do, instead of 1 (above it looks like three of them, but exp(t D) is diagonal so it is in fact a vector-vector product, term by term).
on the other hand, you still need to diagonalize the matrix.
this trick is useful when you have many matrix/length configurations. Even with one single matrix, if you have a continuous gamma distribution or a dirichlet process for describing rates across sites, instead of a discretized gamma distribution, then this is very useful: because, then, you don’t have to cope with the fact that you basically have a different exp(t Q) for each branch and for each site (or each category of the dirichlet process).
so, “technically”, for me, the main contribution of Irvahn and @vminin is not so much “exponentiation-free” than “diagonalization-free” stochastic mapping-- which is also very nice, though. I like very much the idea.

One strategy for jointly updating topologies and augmented substitution histories that Jan and I discussed was to apply a topology rearrangement (e.g., SPR move) and just transfer all virtual and real jump times on the branches when removing and reattaching them. Then, we can apply our MCMC kernel and accept/reject using the likelihood of the complete data – p(V,W) – in our notation. Given enough virtual jumps, this proposal should allow the augmented history to adapt quickly to a new topology. This is of course just a conjecture at this point.

At some point, we wanted to jointly update topologies and substitution histories (when we were implementing structurally-constained, site-interdependent , models of protein sequence evolution with Nicolas Rodrigue).
What I had then in mind was to essentially propose a SPR or NNI, then erase the mapping locally, around the three branches surrounding the regrafting point, and resample it conditional on the end-points.
did you think about doing something along those lines ?

To ensure the sampled histories remain consistent for a prune-regraft proposal, my sense is you should minimally need to update the “start” state of the pruned lineage to match the sampled history at the regraft position, and then resample the history for only that lineage. Resampling the three branches around the regrafting position should improve acceptance probabilities, though you may also want to resample the two branch histories incident to the pruning site (since fewer substitutions may be needed to explain the evolution along that lineage without the pruned clade).
This makes 1, 3, or 5 path samplings used per topology update. But you can also imagine there is are local neighborhoods of branch histories (perhaps by # edges or sum of branch lengths) surrounding the prune and regraft positions that also need to be resampled for good acceptance probabilities.
In any case, fun to think about!

and what would that give if you essentially integrate the likelihood over V over the entire tree, conditional on the numbers of jumps (both real and virtual) along branches (which you can do, as you say in the article, by just using the B^{m_i} for the transition probability matrix along branch i) – and then use this semi-integrated likelihood for a Metropolis-Hastings update in the space of tree topologies ?
or perhaps this is what you are proposing?

Collapsing/integrating over V is an interesting idea. I was thinking about drawing V from p(V | W, \tau^{new}), where \tau^{new} is the proposed phylogeny, and then accepting/rejecting using complete-data likelihood (product of exponentials and such).
@nicolas_lartill, @mlandis, are you interested in collaborating on this? It would be super awesome, I think!

Very good point, @nicolas_lartill. We haven’t used this trick in our comparisons, but we also pretended that we had Q diagonalized to avoid dealing with numerical issues during diagonalization.
I think the only really fair way to compare our new stochastic mapping with the state-of-the-art is to incorporate it into PhyloBayes, RevBayes, or BEAST and see how much it helps when approximating the posterior of phylogenies and other parameters. But it was too much for the first paper, where we just wanted to outline the basic algorithm.

Yeah, I wasn’t too happy about this either. I guess we did this, because plot 3 is the only plot with no virtual jumps, so it is not as busy as the other 3.

oh yes, sure, I understand.
and anyway, there are many other good reasons than just computational gains to develop mapping-based inference.

in fact, I think that these two approaches turn out to be exactly equivalent.
If you do what you say, you will end up simplifying the p(V | W, tau) factors between the ratio of probabilities and the hastings ratio. And thus, everything looks like you are in fact working on the collapsed parameter vector (in which V has been integrated away).
More generally, you can collapse and re-augment your parameter vector at will, depending on the context: you can discard V if you want to make a long series of topological updates, and then resurrect it whenever you need it, by drawing it from the current conditional distribution.
you should of course feel free to use all of those ideas in your future work, if you think they can be useful. I will be happy enough to know that this has been helpful.

This is very kind of you, @nicolas_lartill, but we are not competent enough to do it on our own  Seriously though, Jan and I need to decide which platform to use for going forward with this. PhyloBayes was our first though, but we are afraid to mess with it without someone holding our hands.
Seriously though, Jan and I need to decide which platform to use for going forward with this. PhyloBayes was our first though, but we are afraid to mess with it without someone holding our hands.

I haven’t thought about local proposals, but my feeling, in line with @nicolas_lartill’s thoughts, is that even local rearrangements of phylogenies are better to be accompanied by global updates of the substitution history.

We could certainly think about it. The code of the old version of phylobayes is close to inextricable. On the other hand, the mpi version is a bit more rationalized (although still not totally compliant with current object-oriented programming practice), so it could be used as a possible environment.

in fact, once you realize that resampling V ~ p(V | W, tau) or discarding the V component while performing topological updates are mcmc-equivalent, as I mentioned above, and that the usual caching of conditional likelihoods at the nodes of the tree becomes available in the latter case, then all this means that you can resample the topology in a computationally efficient manner while integrating over all configurations of the V component.

I moved 3 posts to an existing topic: Matrix strategies to avoid rate matrix exponentiation

This is good to know, @nicolas_lartill. I didn’t know about the mpi version of PhyloBayes is so different from the old version.
@mlandis, you mentioned that you plan to work on this using RevBayes as a platform – any thoughts on PhyloBayes vs RevBayes as a platform for moving forward? We wouldn’t want to steal your thunder, but would be happy to help of course. There are plenty of projects to go around in terms of ejecting matrix exponentiation/diagonalization from various phylogenetic models.

I get you point, @nicolas_lartill – clever! Although this is all conditional on the number and locations of real and virtual jumps – I don’t have a sense of how frequently they would have to be updated, but you are probably right in that one does not have to update W immediately following a topology update.

We use the branch indices to map the figure onto our notation in text:
The tree in plot 4 of Figure 1 has four branches so the collection of state labels is V = {v1,v2,v3,v4}, where v1 = (1, 1), v2 = (1, 2), v3 = (3, 3), and v4 = (3, 1). The collection of intertransition times is W = {w1, w2, w3, w4}, where w1 = (1.6, 1.6), w2 = (0.64, 2.56), w3 = (7, 1), and w4 = (2.4,2.4). The locations/times of each virtual jump on branch i is represented by a vector ui = (ui1, …, ui(mi−ni)). For example, the distance from the parent node of branch i to the dth virtual jump is uid. The collection of the ui vectors, fully determined by (V,W), is denoted by U. In plot 4 of Figure 1 the collection of virtual jump times is U = {u1, u2, u3, u4} whereu1 =(1.6),u2 =(),u3 =(7),andu4 =().
It is pretty sad that Jan and I were working on this for more than 2 years (we noticed Rao and Teh’s original paper almost immediately as it came out in print), but still cannot explain the method well.

Ha, I didn’t mean to make you feel badly! I just thought for a while that the numbers were telling me something special about panel 3. I suggest just having a separate panel with the branch numberings.

A separate uncolored tree with branch numbers is a great idea. Thanks, @ematsen!
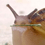
This is great! In addition to improving from O(N^3) to O(N^2) for dense matrices and even to O(N) for sparse enough matrices, the continuous time Bayesian network (CTBN) section in a later section of the Rao-Teh paper (perhaps not implemented here) will allow O(<< N) stochastic mapping with structured (e.g. covarion-like) continuous time Markov processes with combinatorially large state spaces. We are currently looking at a process with N = 61*2^20 ~ 60M states!

Jan and I did not touch CTBN, Markov blankets, etc., so this is not implemented in the paper above. It is great that you find these tools useful for your enormous state space, @argriffing! Soon enough we will be modeling evolution on the state space, where each element is the whole DNA sequence!

Of course there’s no stolen thunder when brainstorming! Either framework suits me, though I’d prefer RevBayes if only for my familiarity with its code. Since you mentioned interest in future projects, I also think it provides a flexible framework for exploring new models (not to imply otherwise of PhyloBayes). In any case, I anticipate identifying good MCMC proposals will be a greater challenge than the software implementation.